I campioni di acqua oceanica producono un tesoro di dati sui virus a RNA
I campioni di acqua oceanica raccolti da tutto il mondo hanno prodotto una miniera di nuovi dati al riguardo[{” attribute=””>RNA viruses, expanding ecological research possibilities and reshaping our understanding of how these small but significant submicroscopic particles evolved.
Combining machine-learning analyses with traditional evolutionary trees, an international team of researchers has identified 5,500 new RNA virus species that represent all five known RNA virus phyla and suggest there are at least five new RNA virus phyla needed to capture them.
The most abundant collection of newly identified species belong to a proposed phylum researchers named Taraviricota, a nod to the source of the 35,000 water samples that enabled the analysis: the Tara Oceans Consortium, an ongoing global study onboard the schooner Tara of the impact of climate change on the world’s oceans.
“There’s so much new diversity here – and an entire phylum, the Taraviricota, were found all over the oceans, which suggests they’re ecologically important,” said lead author Matthew Sullivan, professor of microbiology at The Ohio State University.
“RNA viruses are clearly important in our world, but we usually only study a tiny slice of them – the few hundred that harm humans, plants, and animals. We wanted to systematically study them on a very big scale and explore an environment no one had looked at deeply, and we got lucky because virtually every species was new, and many were really new.”
The study was published on April 7, 2022, in the journal Science.
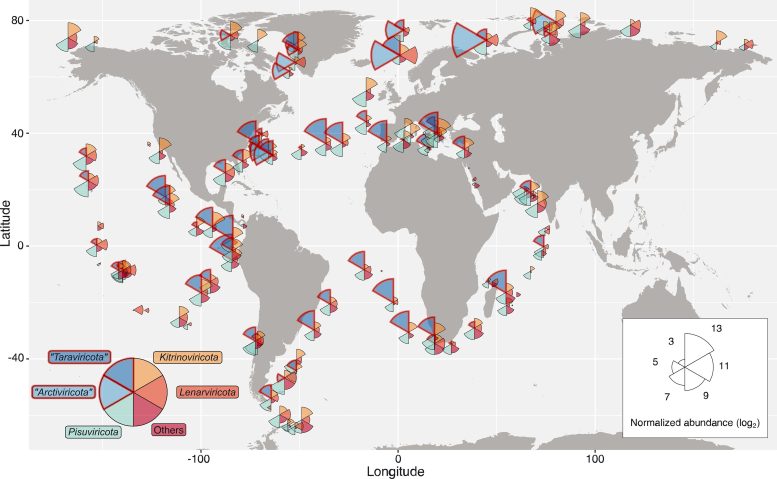
This map shows the distribution of RNA viruses across the ocean. Wedge size is proportional to the average abundance of viruses present in that area, and wedge color indicates virus phyla. Credit: Reprinted with permission from Zayed et al., Science Volume 376:156(2022)
While microbes are essential contributors to all life on the planet, viruses that infect or interact with them have a variety of influences on microbial functions. These types of viruses are believed to have three main functions: killing cells, changing how infected cells manage energy, and transferring genes from one host to another.
Knowing more about virus diversity and abundance in the world’s oceans will help explain marine microbes’ role in ocean adaptation to climate change, the researchers say. Oceans absorb half of the human-generated carbon dioxide from the atmosphere, and previous research by this group has suggested that marine viruses are the “knob” on a biological pump affecting how carbon in the ocean is stored.
By taking on the challenge of classifying RNA viruses, the team entered waters still rippling from earlier taxonomy categorization efforts that focused mostly on RNA viral pathogens. Within the biological kingdom Orthornavirae, five phyla were recently recognized by the International Committee on Taxonomy of Viruses (ICTV).
“RdRp is supposed to be one of the most ancient genes – it existed before there was a need for DNA. So we’re not just tracing the origins of viruses, but also tracing the origins of life.” — Ahmed Zayed
Though the research team identified hundreds of new RNA virus species that fit into those existing divisions, their analysis identified thousands more species that they clustered into five new proposed phyla: Taraviricota, Pomiviricota, Paraxenoviricota, Wamoviricota and Arctiviricota, which, like Taraviricota, features highly abundant species – at least in climate-critical Arctic Ocean waters, the area of the world where warming conditions wreak the most havoc.
Sullivan’s team has long cataloged DNA virus species in the oceans, growing the numbers from a few thousand in 2015 and 2016 to 200,000 in 2019. For those studies, scientists had access to viral particles to complete the analysis.
In these current efforts to detect RNA viruses, there were no viral particles to study. Instead, researchers extracted sequences from genes expressed in organisms floating in the sea, and narrowed the analysis to RNA sequences that contained a signature gene, called RdRp, which has evolved for billions of years in RNA viruses, and is absent from other viruses or cells.
Because RdRp’s existence dates to when life was first detected on Earth, its sequence position has diverged many times, meaning traditional phylogenetic tree relationships were impossible to describe with sequences alone. Instead, the team used machine learning to organize 44,000 new sequences in a way that could handle these billions of years of sequence divergence, and validated the method by showing the technique could accurately classify sequences of RNA viruses already identified.
“We had to benchmark the known to study the unknown,” said Sullivan, also a professor of civil, environmental and geodetic engineering, founding director of Ohio State’s Center of Microbiome Science and a leadership team member in the EMERGE Biology Integration Institute.
“We’ve created a computationally reproducible way to align those sequences to where we can be more confident that we are aligning positions that accurately reflect evolution.”
Further analysis using 3D representations of sequence structures and alignment revealed that the cluster of 5,500 new species didn’t fit into the five existing phyla of RNA viruses categorized in the Orthornavirae kingdom.
“We benchmarked our clusters against established, recognized phylogeny-based taxa, and that is how we found we have more clusters than those that existed,” said co-first author Ahmed Zayed, a research scientist in microbiology at Ohio State and a research lead in the EMERGE Institute.
In all, the findings led the researchers to propose not only the five new phyla, but also at least 11 new orthornaviran classes of RNA viruses. The team is preparing a proposal to request formalization of the candidate phyla and classes by the ICTV.
Zayed said the extent of new data on the RdRp gene’s divergence over time leads to a better understanding about how early life may have evolved on the planet.
“RdRp is supposed to be one of the most ancient genes – it existed before there was a need for DNA,” he said. “So we’re not just tracing the origins of viruses, but also tracing the origins of life.”
Reference: “Cryptic and abundant marine viruses at the evolutionary origins of Earth’s RNA virome” by Ahmed A. Zayed, James M. Wainaina, Guillermo Dominguez-Huerta, Eric Pelletier, Jiarong Guo, Mohamed Mohssen, Funing Tian, Akbar Adjie Pratama, Benjamin Bolduc, Olivier Zablocki, Dylan Cronin, Lindsey Solden, Erwan Delage, Adriana Alberti, Jean-Marc Aury, Quentin Carradec, Corinne da Silva, Karine Labadie, Julie Poulain, Hans-Joachim Ruscheweyh, Guillem Salazar, Elan Shatoff, Tara Oceans Coordinators, Ralf Bundschuh, Kurt Fredrick, Laura S. Kubatko, Samuel Chaffron, Alexander I. Culley, Shinichi Sunagawa, Jens H. Kuhn, Patrick Wincker, Matthew B. Sullivan, Silvia G. Acinas, Marcel Babin, Peer Bork, Emmanuel Boss, Chris Bowler, Guy Cochrane, Colomban de Vargas, Gabriel Gorsky, Lionel Guidi, Nigel Grimsley, Pascal Hingamp, Daniele Iudicone, Olivier Jaillon, Stefanie Kandels, Lee Karp-Boss, Eric Karsenti, Fabrice Not, Hiroyuki Ogata, Nicole Poulton, Stéphane Pesant, Christian Sardet, Sabrinia Speich, Lars Stemmann, Matthew B. Sullivan, Shinichi Sungawa and Patrick Wincker, 7 April 2022, Science.
DOI: 10.1126/science.abm5847
This research was supported by the National Science Foundation, the Gordon and Betty Moore Foundation, the Ohio Supercomputer Center, Ohio State’s Center of Microbiome Science, the EMERGE Biology Integration Institute, the Ramon-Areces Foundation and Laulima Government Solutions/NIAID. The work was also made possible by the unprecedented sampling and science of the Tara Oceans Consortium, the nonprofit Tara Ocean Foundation and its partners.
Additional co-authors on the paper were co-lead authors James Wainaina and Guillermo Dominguez-Huerta, as well as Jiarong Guo, Mohamed Mohssen, Funing Tian, Adjie Pratama, Ben Bolduc, Olivier Zablocki, Dylan Cronin and Lindsay Solden, all of Sullivan’s lab; Ralf Bundschuh, Kurt Fredrick, Laura Kubatko and Elan Shatoff of Ohio State’s College of Arts and Sciences; Hans-Joachim Ruscheweyh, Guillem Salazar and Shinichi Sunagawa of the Institute of Microbiology and Swiss Institute of Bioinformatics; Jens Kuhn of the National Institute of Allergy and Infectious Diseases; Alexander Culley of the Université Laval; Erwan Delage and Samuel Chaffron of the Université de Nantes; and Eric Pelletier, Adriana Alberti, Jean-Marc Aury, Quentin Carradec, Corinne da Silva, Karine Labadie, Julie Poulain and Patrick Wincker of Genoscope.


